Last Updated: May 29, 2026
Introduction to Growth Factors
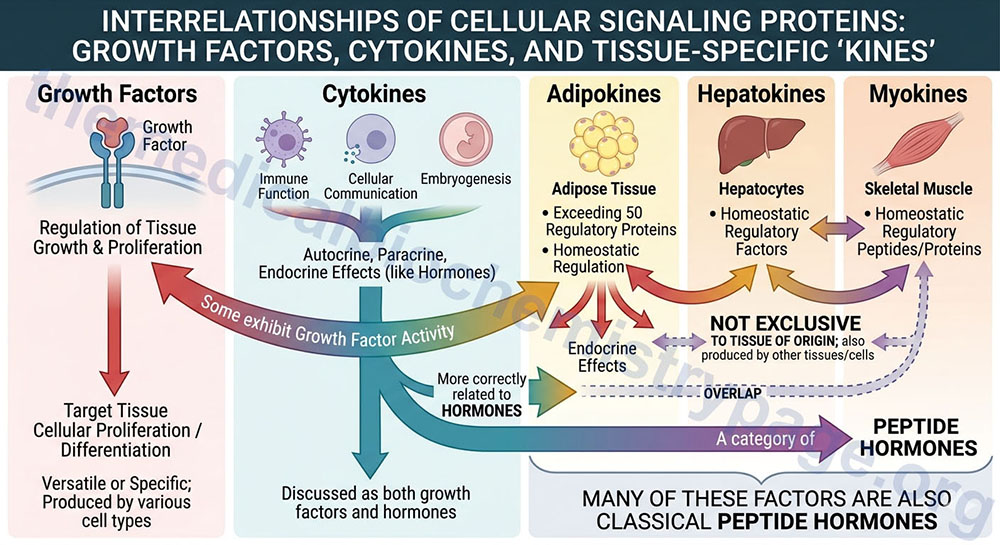
The term growth factor is generally used to describe a protein or peptide whose function is predominantly, although not exclusively, related to the regulation of target tissue growth and potential proliferation. Growth factors are, therefore, proteins or peptides that are produced by a variety of different cell types and when released into the vasculature interact with and bind to specific receptors on the cell surface eliciting responses within the target tissue. The primary result of activating growth factor receptors is cellular proliferation and/or differentiation. Many growth factors are quite versatile, stimulating not only cell growth but also cellular division in numerous different cell types, while others are specific to a particular cell-type.
Cytokines are a class of signaling proteins that are used extensively in cellular communication, immune function and embryogenesis. Cytokines are produced by a variety of hematopoietic and non-hematopoietic cell types and can exert autocrine, paracrine and endocrine effects as do the hormones. They are, therefore, more correctly related to hormones than to growth factors in their overall functions. However, many cytokines also exhibit growth factor activity so they are discussed here as well as in the Peptide Hormones page.
Numerous tissues produce and secrete peptides and proteins that exert effects that, in some cases are growth stimulating, and in others are metabolic and/or cellular homeostasis regulating. Using the terminology adopted for hematopoietic modulating proteins (the cytokines) the descriptors for these peptides and proteins includes “kine” as the suffix. Adipose tissue secretes in excess of 50 different homeostatic regulatory proteins and these factors are collectively termed adipokines. Similarly the liver, primarily hepatocytes, also secretes a number of homeostatic regulatory factors and they are collectively referred to as hepatokines. Skeletal muscle also synthesizes and secretes several homeostatic regulatory peptides and proteins termed myokines. Although many of these peptides and proteins are exclusive to a specific tissues many of the adipokines, hepatokines, and myokines are not exclusive to these tissues but are produced by several other tissues and cells. Many of these factors are also classical peptide hormones and more details are discussed in the Peptide Hormones and Their Receptors page.
The lists in the following Tables, as well as the descriptions of several factors, are not intended to be comprehensive nor complete but a look at some of the more commonly known factors and their principal activities.
Table of Representative Growth Factors
| Factor | Principal Source | Primary Activity | Comments |
| EGF | submaxillary gland, Brunners gland | promotes proliferation of mesenchymal, glial and epithelial cells | represents the founding member of the EGF-family of proteins that includes, but is not limited to, transforming growth factor-α (TGF-α), amphiregulin, and the neuregulins (neuregulin-1, -2, -3, and -4) |
| Erythropoietin | kidney | promotes proliferation and differentiation of erythrocytes | |
| FGF | wide range of cells; protein is associated with the ECM | promotes proliferation of many cells; inhibits some stem cells; induces mesoderm to form in early embryos | at least 18 family members, 5 distinct receptors |
| IGF-1 | primarily liver | promotes proliferation of many cell types | related to IGF-2 and proinsulin, also called somatomedin C |
| IGF-2 | variety of cells | promotes proliferation of many cell types primarily of fetal origin | related to IGF-1 and proinsulin |
| NGF | mast cells, eosinophils, bone marrow stromal cells, keratinocytes | promotes neurite outgrowth and neural cell survival | member of a family of proteins termed neurotrophins that promote proliferation and survival of neurons; neurotrophin receptors are a class of related proteins first identified as proto-oncogenes: TrkA (“trackA”), TrkB, TrkC |
| PDGF | platelets, endothelial cells, placenta | promotes proliferation of connective tissue, glial and smooth muscle cells | represents a family of four peptides encoded by four distinct genes: A, B, C, and D; these four peptides form either homo- or heterodimers such that five distinct biologically active PDGF isoforms (AA, AB, BB, CC, DD) result |
| TGF-α | macrophages, keratinocytes, hypothalamic astrocytes; commonly expressed by transformed cells | important for normal wound healing, cellular proliferation, female reproductive maturation, embryogenesis | is a member of the EGF-family of proteins; functions by binding to the EGF receptor |
| TGF-β | activated Th1 cells (T-helper) and natural killer (NK) cells | anti-inflammatory (suppresses cytokine production and class II MHC expression), promotes wound healing, inhibits macrophage and lymphocyte proliferation | at least 100 different family members |
Interleukins and Cytokines
Cytokines are a unique family of growth factors. Secreted primarily from leukocytes, cytokines stimulate both the humoral and cellular immune responses, as well as the activation of phagocytic cells. Cytokines that are secreted from lymphocytes are termed lymphokines, whereas those secreted by monocytes or macrophages are termed monokines. A large family of cytokines are produced by various cells of the body. Many of the lymphokines are also known as interleukins (ILs), since they are not only secreted by leukocytes but also able to affect the cellular responses of leukocytes. Specifically, interleukins are growth factors targeted to cells of hematopoietic origin. The list of identified interleukins grows continuously with the total number of individual activities now at 43 (18 are listed in the following Table).
Table of Representative Interleukins and Interferons
| Interleukins | Principal Source | Primary Activity |
| IL1-α and -β | macrophages and other antigen presenting cells (APCs) | co-stimulation of APCs and T cells, inflammation and fever, acute phase response, hematopoiesis |
| IL-2 | activated Th1 cells, NK cells | proliferation of B cells and activated T cells, NK functions |
| IL-3 | activated T cells | growth of hematopoietic progenitor cells |
| IL-4 | Th2 and mast cells | B cell proliferation, eosinophil and mast cell growth and function, IgE and class II MHC expression on B cells, inhibition of monokine production |
| IL-5 | Th2 and mast cells | eosinophil growth and function |
| IL-6 | activated Th2 cells, APCs, other somatic cells such as hepatocytes and adipocytes | acute phase response, B cell proliferation, thrombopoiesis, synergistic with IL-1β and TNF on T cells |
| IL-7 | thymic and marrow stromal cells | T and B lymphopoiesis |
| IL-8 | macrophages, other somatic cells | chemoattractant for neutrophils and T cells |
| IL-9 | T cells | hematopoietic and thymopoietic effects |
| IL-10 | activated Th2 cells, CD8+ T and B cells, macrophages | inhibits cytokine production, promotes B cell proliferation and antibody production, suppresses cellular immunity, mast cell growth |
| IL-11 | bone marrow stromal cells | synergistic hematopoietic and thrombopoietic effects |
| IL-12 | B cells, T cells, macrophages, dendritic cells | proliferation of NK cells, INF-γ production, promotes cell-mediated immune functions |
| IL-13 | Th2 cells, B cells, macrophages | stimulates growth and proliferation of B cells, inhibits production of macrophage inflammatory cytokines |
| IL-14 | T cells and malignant B cells | regulates the growth and proliferation of B cells |
| IL-15 | monocytes, thyroid, lymph nodes, myocytes | induces production of NK cells |
| IL-16 | eosinophils, CD8+ T cells, lymphocytes, epithelial cells | chemoattractant for CD4+ cells |
| IL-17: six isoforms all from different genes; IL-17A, B, C, D, E, and F (IL-17E also called IL-25) | A and F forms only expressed in a subset of T cells; B expressed in leukocytes and peripheral tissues; C up-regulated during inflammation; D expressed in nervous system and skeletal muscle; E expressed in peripheral tissues | increases production of inflammatory cytokines, angiogenesis, affects endothelial and epithelial cells |
| IL-18 | macrophages | increases NK cell activity, induces production of INF-γ |
| Interferons | Principal Source | Primary Activity |
| INF-α and -β | macrophages, neutrophils and some somatic cells | antiviral effects, induction of class I MHC on all somatic cells, activation of NK cells and macrophages |
| INF-γ | activated Th1 and NK cells | induces of class I MHC on all somatic cells, induces class II MHC on APCs and somatic cells, activates macrophages, neutrophils, NK cells, promotes cell-mediated immunity, antiviral effects |
Adipose Tissue Hormones and Adipokines
Adipose tissue is not merely an organ designed to passively store excess carbon in the form of fatty acids esterified to glycerol (triglycerides). Mature adipocytes synthesize and secrete numerous enzymes, growth factors, cytokines and hormones that are involved in overall energy homeostasis. Many of the factors that influence adipogenesis are also involved in diverse processes in the body including lipid homeostasis and modulation of inflammatory responses. In addition, a number of proteins secreted by adipocytes play important roles in these same processes. In fact, recent evidence has demonstrated that many factors secreted from adipocytes are proinflammatory mediators and these proteins have been termed adipocytokines or adipokines. Members of this class of protein secreted from adipocytes include TNFα, IL-6 and leptin.
For descriptions of many of the identified adipokines go to the Secreted Factors: Tissue Kines page.
Liver Derived Hepatokines
Hepatocytes produce and secrete hundreds of proteins and peptides. Indeed, a major function of the liver is to produce a multitude of proteins that are found circulating in the vasculature and perform a wide array of functions. Many of the factors secreted by hepatocytes are defined as hepatokines and when released from the liver they exert global whole body homeostatic processes. The major pathophysiologically significant hepatokines are those that exert regulatory effects on overall lipoid and glucose homeostasis. As might be expected, disruptions in the regulated synthesis and release of these hepatokines contribute to the pathologies of obesity and type 2 diabetes.
For descriptions of many of the identified hepatokines go to the Secreted Factors: Tissue Kines page.
Skeletal Muscle Derived Myokines
Like adipose tissue and the liver, skeletal muscle produces and secretes a number of proteins, peptides, and metabolites that exert autocrine, paracrine, and endocrine effects. These secreted substances are collectively referred to as myokines. Like adipokines and hepatokines, many of the regulatory substances secreted from skeletal muscle are also secreted by other tissues such as the liver and adipose tissue.
For descriptions of many of the identified myokines go to the Secreted Factors: Tissue Kines page.
Exerkines: Exercise-Induced Kines
For descriptions of many of the identified exerkines go to the Secreted Factors: Tissue Kines page.
Epidermal Growth Factor (EGF)
EGF is synthesized as a preproprotein that is processed to a 53 amino acid functional growth factor. The EGF preproprotein is derived from the EGF gene. The EGF gene is located on chromosome 4q25 and is composed of 24 exons that generate four alternatively spliced mRNAs, each of which encode distinct protein isoforms.
There are several proteins that are related to EGF in that they bind and activate the same receptors. The EGF family of proteins includes TGF-α, amphiregulin, epiregulin, heparin-binding EGF-like growth factor (HB-EGF), epigen (epithelial mitogen), betacellulin, and the four neuregulin proteins.
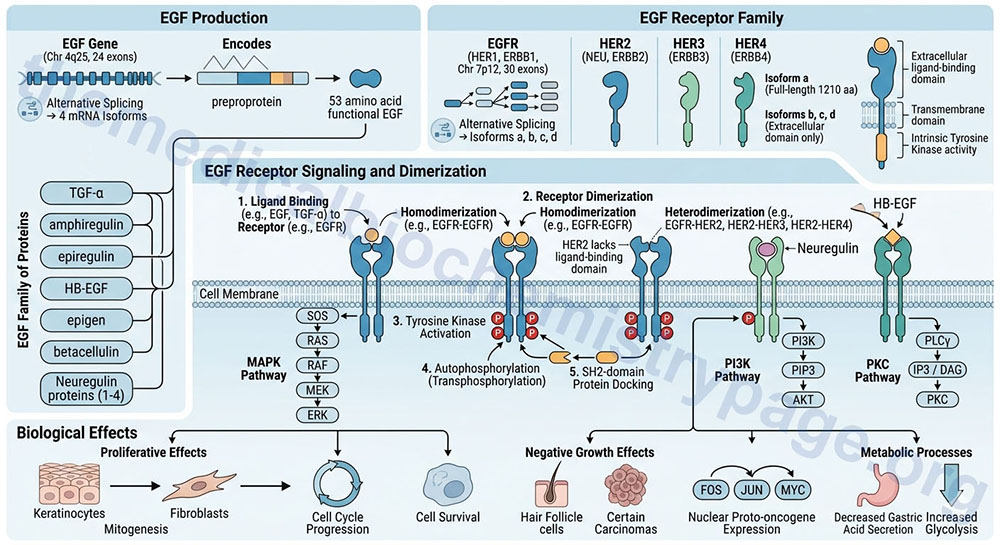
Like all growth factors, EGF binds to specific high-affinity, low-capacity receptors (EGFR) on the surface of responsive cells. Intrinsic to the EGF receptor is tyrosine kinase activity, which is activated in response to EGF binding and receptor dimerization. In the absence of ligand the EGFR exists as a monomer in the membrane. The kinase domain of the EGF receptor phosphorylates the EGF receptor itself (autophosphorylation; also referred to as transphosphorylation). These phosphotyrosine residues act as docking sites for numerous signal transduction proteins that contain an SH2 domain.
The EGF receptor is derived from the EGFR gene. The EGFR gene is located on chromosome 7p12 and is composed of 30 exons that generate four alternatively spliced mRNAs, each of which encode a distinct protein. The major transmembrane-spanning EGF receptor is derived from the EGFR isoform a encoding mRNA. This precursor protein is composed of 1210 amino acids. The EGFR isoforms b, c, and d mRNAs encoded proteins that only contain the extracellular domain of the full-length isoform a receptor. The EGFR gene encoded protein is also, more commonly referred to as HER1, for human EGF receptor 1. The EGFR protein is also known as ERBB1 since it is a member of the ErbB family of receptor tyrosine kinases.
There are three additional EGF receptor family proteins that are encoded by the ERBB2, ERBB3, and ERBB4 genes. These designations refer to their initial identification as being receptors for the ERB-B oncogene encoded by the avian erythroblastosis virus. The ERBB2 gene encoded protein is most commonly referred to as HER2 but is also known as the NEU proto-oncogene. The ERBB3 gene encoded protein is also known as HER3 and the ERBB4 gene encoded protein is also referred to as HER4.
All of the EGF receptor family members, except HER2, dimerize in response to ligand binding. Since the ERBB2 encoded protein (HER2) lacks a ligand-binding domain it does not dimerize in response to ligand binding. In addition to EGF the EGFR encoded protein is known to bind several members of the EGF family of proteins that includes TGF-α, amphiregulin, epiregulin, epigen, HB-EGF, and betacellulin. The ERBB3 encoded protein is the receptor for neuregulin 1 and neuregulin 2. The ERBB4 encode protein binds HB-EGF and all four of the neuregulin proteins. Upon ligand binding the EGFR encoded protein forms homodimers but can also form heterodimers with the ERBB2 encoded receptor. The ERBB2 encoded receptor also forms heterodimers with the ERBB3 and ERBB4 encoded receptors.
EGF has proliferative effects on cells of both mesodermal and ectodermal origin, particularly keratinocytes and fibroblasts. EGF exhibits negative growth effects on certain carcinomas as well as hair follicle cells. Growth-related responses to EGF include the induction of nuclear proto-oncogene expression, such as FOS, JUN and MYC. EGF also exerts effects on metabolic processes such as decreasing gastric acid secretion, and increasing the rate of glycolysis.
The growth and proliferative effects of EGF as well as the metabolic effects are exerted in response to the activation of numerous divergent signal transduction pathways in response to EGF-mediated activation of the EGFR. The activated EGF receptor family members trigger activation of mitogen-activated protein kinase (MAPK), phosphatidylinositol-4,5-bisphosphate 3-kinase (PI3K), and protein kinase C (PKC). The primary effects of EGF receptor signaling are cell cycle progression, cell survival, and cell proliferation.
HER2 and Breast Cancer
Unlike the other EGF receptor family members, the ERBB2 encoded protein (HER2) does not bind ligand due to the lack of a ligand-binding domain. The conformation of HER2 in the membrane closely mimics the ligand-bound EGFR, ERBB3, and ERBB4 encoded proteins, i.e. it is likely to be constitutively in the activated receptor homodimer state. In addition, HER2 interacts with each of the other ligand-bound EGFR family members forming heterodimers. The formation of these heterodimeric complexes results in enhancement of the receptor tyrosine kinase-mediated downstream signal transduction pathways.
Amplification and/or overexpression of the ERBB2 gene is associated with numerous cancers including in 15–30% of breast cancers and 10–30% of gastric/gastroesophageal cancers. Overexpression of the ERBB2 gene has also been seen in other cancers like ovary, endometrium, bladder, lung, colon, and head and neck. As such, the level of ERBB2 expression serves as a prognostic and predictive cancer biomarker.
HER2-positive breast cancers may have up to 25–50 copies of the ERBB2 gene resulting in a 40- to 100-fold increase in HER2 protein. The amplification of the ERBB2 gene in breast cancer is associated with shorter disease-free states as well as reduced overall survival rates. In some breast cancers there is an aberrant form of HER2 (known as p95) which lacks the extracellular domain. The p95 form of HER2 is constitutively active and results in breast cancers that are resistance to trastuzumab (Herceptin™). Herceptin is a monoclonal antibody used in the treatment of HER2 positive breast cancers but is also used in the treatment of some forms of stomach cancer that are HER2 positive.
Platelet-Derived Growth Factor (PDGF)
The PDGF is either a homodimeric or heterodimeric growth factor. The PDGF composition is determined by the expression of four distinct polypeptides encoded by four different genes. The PDGF peptides are identified as PDGF-A, -B, -C, and -D. These four PDGF peptides result in five distinct dimeric forms of PDGF (PDGF-AA, -AB, -BB, -CC, and -DD). The SIS proto-oncogene has been shown to be homologous to the PDGF-B peptide. Only dimeric forms of PDGF interact with the PDGF receptors.
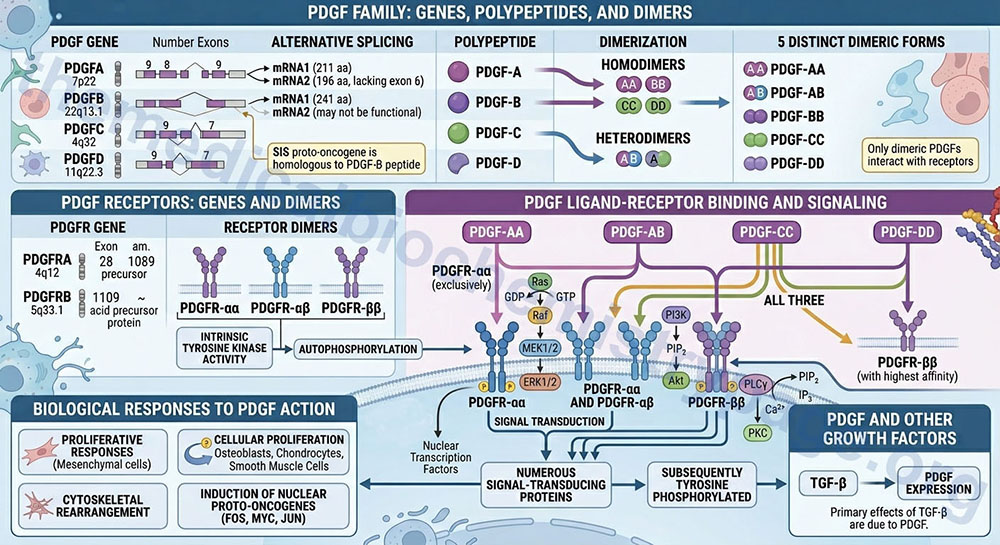
The PDGF-A preproprotein is derived from the PDGFA gene. The PDGFA gene is located on chromosome 7p22 and is composed of 9 exons that generate two alternatively spliced mRNAs. PDGF-A isoform 1 is a 211 amino acid preproprotein and isoform 2, which lacks the coding information from exon 6, is a 196 amino acid preproprotein.
The PDGF-B preproprotein is derived from the PDGFB gene. The PDGFB gene is located on chromosome 22q13.1 and is composed of 8 exons that generate two alternatively spliced mRNAs. PDGF-B isoform 1 is a 241 amino acid preproprotein. The PDGF-B isoform 2 protein may not undergo processing to a function protein.
The PDGF-C preproprotein is derived from the PDGFC gene. The PDGFC gene is located on chromosome 4q32 and is composed of 9 exons that encode a 345 amino acid preproprotein.
The PDGF-D preproprotein is derived from the PDGFD gene. The PDGFD gene is located on chromosome 11q22.3 and is composed of 7 exons that generate two alternatively spliced mRNAs. PDGF-D isoform 1 is a 370 amino acid preproprotein and PDGF-D isoform 2 is a 364 amino acid preproprotein.
Three distinct forms of the PDGF receptor have been identified that result from the dimerization of proteins expressed from two different genes. The composition of these three receptor types are αα, αβ, and ββ. Like the EGF receptor, the PDGF receptors have intrinsic tyrosine kinase activity. Following autophosphorylation of the PDGF receptor, numerous signal-transducing proteins associate with the receptor and are subsequently tyrosine phosphorylated.
The PDGF receptor α (alpha) protein is encoded by the PDGFRA gene. The PDGFRA gene is located on chromosome 4q12 and is composed of 28 exons that encode a 1089 amino acid precursor protein.
The PDGF receptor β (beta) protein is encoded by the PDGFRB gene. The PDGFRB gene is located on chromosome 5q33.1 and is composed of 26 exons that encode a amino acid precursor protein.
The PDGF-AA isoform binds exclusively to the PDGFR-αα type receptor. The PDGF-BB isoform can bind to all three types for PDGFR. The PDGF-AB isoform binds to the PDGFR-αα and PDGFR-αβ type receptors. The PDGF-CC isoform, like the PDGF-AB isoforms, specifically binds to the PDGFR-αα and PDGFR-αβ type receptors. The PDGF-DD isoform binds with highest affinity to the PDGFR-ββ type receptors.
Proliferative responses to PDGF action are exerted on many mesenchymal cell types. Other growth-related responses to PDGF include cytoskeletal rearrangement and increased polyphosphoinositol turnover. Again, like EGF, PDGF induces the expression of a number of nuclear localized proto-oncogenes, such as FOS, MYC, and JUN. Indeed, the primary effects of TGF-β are due to the induction, by TGF-β, of PDGF expression.
Fibroblast Growth Factors (FGFs)
The fibroblast growth factor (FGF) family consists of 22 genes in the human genome. Of these 22 genes there are currently 18 that produce growth factors that have been characterized to function through activation of the FGF receptors. These members are numbered FGF1–FGF10 and FGF16–FGF23. These 18 proteins are divided into six different FGF families based upon differences in sequence homology.
- Family 1 (FGF1 subfamily): FGF1 and FGF2
- Family 2 (FGF7 subfamily): FGF3, FGF7, FGF10, and FGF22
- Family 3 (FGF4 subfamily): FGF4, FGF5, and FGF6
- Family 4 (FGF8 subfamily): FGF8, FGF17, and FGF18
- Family 5 (FGF9 subfamily): FGF9, FGF16, and FGF20
- Family 6 (FGF19 subfamily):FGF19, FGF21, and FGF23
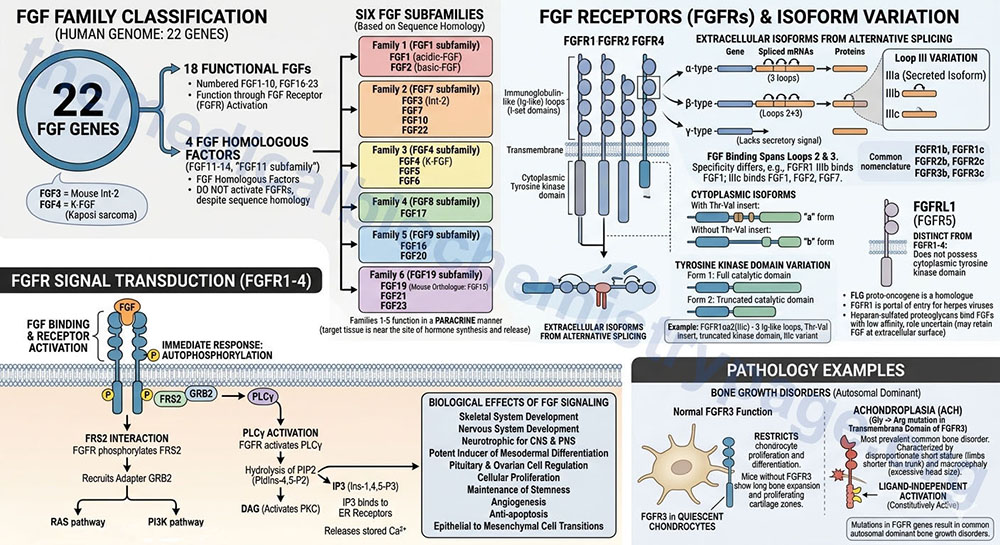
There are four additional genes in humans that express FGF-related proteins (FGF11–FGF14; also referred to as the FGF11 subfamily). Although the FGF11 subfamily proteins contain amino acid sequence homology to members of the six functional FGF subfamilies they do not activate the FGF receptors and are thus, not considered members of the FGF family but are FGF homologous factors. Of note is the fact that human FGF19 is the orthologue of mouse FGF15.
The two originally characterized FGFs were identified by biological assay and are termed FGF1 (acidic-FGF, aFGF) and FGF2 (basic-FGF, bFGF). In mice, the mammary tumor virus integrates at two predominant sites in the mouse genome identified as Int-1 and Int-2. The protein encoded by the Int-2 locus turned out to be a homologue of the FGF family of growth factors and is now called FGF3. Kaposi sarcoma cells (prevalent in patients with AIDS) were found to secrete a homologue of FGF originally called the K-FGF proto-oncogene, it is now known as FGF4.
Studies of human disorders as well as gene knock-out studies in mice show the prominent role for FGFs is in the development of the skeletal system and nervous system in mammals. FGFs also are neurotrophic for cells of both the peripheral and central nervous system. Additionally, several members of the FGF family are potent inducers of mesodermal differentiation in early embryos. Non-proliferative effects include regulation of pituitary and ovarian cell function. The members of the first five families of FGFs all function in a paracrine manner (meaning the target tissue is near the site of hormone synthesis and release).
The FGF Receptors, FGFR
The FGF family member proteins interact with specific cell-surface receptors, FGFR. There are five distinct FGFR encoding genes identified as FGFR1–FGFR4, and FGFRL1, that bind the various FGF ligands. The FGFRL1 protein was originally identified as FGFR5. The FGFR1-FGFR4 proteins belong to the large family of I-set domain containing proteins that as a group are a subfamily of the larger immunoglobulin domain containing superfamily of proteins.
Due to alternative splicing of the mRNAs transcribed from the FGFR1, FGFR2, and FGFR3 genes there are several receptor isoforms produced from both genes. The complexity of FGF receptor isoforms is further compounded by a complex nomenclature system. Each of the FGFR gene encoded proteins can contain three immunoglobulin-like (Ig-like) loops (domains) in the extracellular portion of the protein. As a result of the alternative splicing if the resultant protein contains three Ig-like loops it is referred to as an alpha(α)-type receptor. If the protein possesses only the second and third Ig-like loops it is referred to as a beta(β)-type receptor. If the encoded protein lacks a secretory signal it is referred to as a gamma(γ)-type receptor.
The binding sites for FGF span the second and third Ig-like loops. The third Ig-like loop (identified by the Roman numeral system: III) of both the FGFR1 and FGFR2 encoded proteins exists in three isoforms identified as IIIa, IIIb, and IIIc. The FGFR proteins that contain the IIIa isoform are secreted isoforms. The IIIb and IIIc isoforms of the receptors differ in specificity for different FGF proteins for example the FGFR1 IIIb isoform binds FGF1 while the IIIc form binds FGF1, FGF2, and FGF7.
The nomenclature of the different FGFR isoforms that result from the presence of the IIIb or IIIc Ig loop (domain) is more commonly simplified such that the FGFR1 receptor possessing the IIIb domain is identified as FGFR1b. With this latter nomenclature the FGFR1 gene generates the FGFR1b and FGFR1c isoforms, the FGFR2 gene generates the FGFR2b and FGFR2c isoforms, and the FGFR3 gene generates the FGFR3b and FGFR3c isoforms.
Within the cytoplasmic portion of the receptors if there is a Thr-Val insert, distal to the membrane spanning domain, the receptor is identified as an “a” form. If the Thr-Val insert is absent it is identified as a “b” form.
Finally, with respect to the tyrosine kinase domain, a full catalytic domain receptor is form 1 while a truncated catalytic domain receptor is form 2. As an example, the FGFR1 isoform identified as αa2(IIIc) results from the alternatively spliced mRNA from the FGFR1 gene that encodes a protein containing three Ig-like loops in the extracellular region, a Thr-Val insert after the transmembrane domain, a truncated tyrosine kinase domain, and the IIIc variant of the third Ig-like loop.
Each of the four receptors encoded by the FGFR1–FGFR4 genes has intrinsic tyrosine kinase activity similar to other receptor tyrosine kinase (RTK) family members such as the EGF and PDGF receptors. As with all transmembrane receptors that have tyrosine kinase activity, autophosphorylation of the receptor is the immediate response to FGF binding. Following activation of FGF receptors, numerous signal-transducing proteins associate with the receptor and become tyrosine-phosphorylated.
The primary proteins that interact with phosphorylated tyrosine residues in the various FGFR are FGF receptor substrate 2 (FRS2) and phospholipase C-gamma (PLCγ). When FRS2 is phosphorylated on tyrosine residues by FGFR it recruits the adapter protein growth factor receptor bound protein 2 (GRB2). The interaction of FRS2 with GRB2 then stimulates both the RAS monomeric G-protein signaling pathway and the phosphatidylinositol 3-kinase (PI3K) signaling pathway. The activation of PLCγ by the FGFR results in the hydrolysis of membrane-associated phosphatidylinositol-4,5-bisphosphate (PIP2; PtdIns-4,5-P2) into diacylglycerol (DAG) and inositol-1,4,5-trisphosphate (IP3; Ins-1,4,5-P3). The DAG activates protein kinase C (PKC) and the IP3 binds to receptors on the endoplasmic reticulum stimulating release of stored Ca2+.
The signal transduction cascades activated by FGF receptors stimulate numerous events that includes, but is not limited to, cellular proliferation, maintenance of stemness, angiogenesis, anti-apoptosis, and epithelial to mesenchymal cell transitions.
The additional protein of the FGF receptor family encoded by the FGF receptor like 1 (FGFRL1) gene is distinct from the FGFR1–FGFR4 receptors. The FGFRL1 gene was originally identified as FGFR5, however, unlike the other four FGF receptors the protein encoded by the FGFRL1 gene does not possess a cytoplasmic tyrosine kinase domain.
The FLG proto-oncogene is a homologue of the FGF receptor family. The FGFR1 receptor also has been shown to be the portal of entry into cells for herpes viruses. FGFs also bind to cell-surface heparan-sulfated proteoglycans with low affinity relative to that of the specific receptors. The purpose in binding of FGFs to these proteoglycans is not completely understood but may allow the growth factor to remain associated with the extracellular surface of cells that they are intended to stimulate under various conditions.
The FGF receptors are widely expressed in developing bone and several common autosomal dominant disorders of bone growth have been shown to result from mutations in the FGFR genes. The most prevalent is achondroplasia, ACH. ACH is characterized by disproportionate short stature, where the limbs are shorter than the trunk, and macrocephaly (excessive head size). Almost all persons with ACH exhibit a glycine to arginine substitution in the transmembrane domain of FGFR3. This mutation results in ligand-independent activation of the receptor. FGFR3 is predominantly expressed in quiescent chondrocytes where it is responsible for restricting chondrocyte proliferation and differentiation. In mice with inactivating mutations in FGFR3 there is an expansion of long bone growth and zones of proliferating cartilage further demonstrating that FGFR3 is necessary to control the rate and amount of chondrocyte growth.
The FGF19 Subfamily
The proteins of the sixth FGF subfamily (members FGF19, FGF21, and FGF23) each act in an endocrine manner (meaning the target tissue is distant from the site of hormone synthesis and release) to regulate glucose, cholesterol, bile acid, vitamin D, and phosphate homeostasis. Although FGF19, FGF21, and FGF23 interact with known FGF receptors they do so only in the presence of a binding partner. This binding partner is identified as Klotho (also known as alpha Klotho: αKlotho). The Klotho gene was originally isolated from a mouse model of age-related disorders and thus, the gene was named after the Fate of Greek mythology who spins the thread of life.
Subsequent to the isolation of the αKlotho gene another related gene termed βKlotho was identified. Both αKlotho and βKlotho are involved in the interactions of FGF19, FGF21, and FGF23 with FGF receptors. Although these three FGFs belong to a distinct FGF subfamily, and each acts as an endocrine factor, they have distinct physiological roles.
FGF19 is involved in the control of cholesterol and bile acid synthesis. FGF19 is also involved is the control feeding behaviors through its effects within the hypothalamus. The details of FGF19 function in the regulation of hypothalamic activity are discussed in the Gut-Brain Interrelationships page.
FGF21 exerts numerous biological effects, the details of which are described below.
FGF23 is a bone-derived hormone that plays an important role as a potent regulator of vitamin D function and of phosphate metabolism. The effects of FGF23 are exerted primarily within the kidney where its actions result in increased phosphate excretion (decreased phosphate reabsorption) and suppression of calcitriol (1,25-dihydroxyvitamin D3) synthesis.
Biological Functions of FGF21
Expression of the FGF21 gene is essentially exclusive to the liver although some low level expression is found in pancreas, adipose tissue, and muscle. However, all of the circulating FGF21 is derived from the liver under normal physiological conditions.
Under basal conditions the level of expression of FGF21 is low but expression is significantly enhanced under conditions of nutritional stress as well as other cellular stress conditions. During periods of prolonged fasting or with the consumption of a ketogenic (high-fat low-carbohydrate) diet, expression of the FGF21 gene is induced by peroxisome proliferator activated receptor alpha (PPARα).
In addition to nutrient deprivation, the FGF21 gene is transcriptionally activated by protein restriction and by high glucose intake. The regulation by protein restriction is exerted by activating transcription factor 4 (ATF4) and by nuclear respiratory factor 1 (NRF1). The regulation by sugars (glucose primarily) is exerted by carbohydrate-responsive element-binding protein (ChREBP). These intricate regulatory controls on FGF21 expression ensure that there is a maximal level of FGF21 protein under conditions of a high carbohydrate, low protein diet.
Naturally occurring compounds, that are enriched in a plant-based diet, are also known to result in increased expression of the FGF21 gene. These compounds include the polyphenols of the anthocyanin family, curcumin, and resveratrol.
In addition to transcriptional regulation, the levels of FGF21 protein found in the plasma is regulated by secretion and by proteolytic degradation by enzymes of the serine protease family. The primary proteases that digest FGF21 are dipeptidyl peptidase IV (DPP-IV or DPP4) and fibroblast activation protein alpha (FAP).
FGF21 functions as an autocrine, paracrine, and an endocrine hormone. FGF21 is involved in the regulation of glucose and lipid homeostasis. FGF21 is also important to digestive processes through its actions on exocrine pancreatic cells stimulating them to release their zymogen granule contents into the small intestine.
FGF21 exerts its effects upon binding to specific FGF receptor complexes. These FGF21 receptor complexes consist of an FGF receptor (FGFR) and the co-receptor protein β-Klotho (KLB). FGF21 binds directly to both of these proteins to activate FGFR signaling activity. Studies with cells in culture have found that FGF21 can act through KLB complexed with either the FGFR1c, FGFR2c, or FGFR3c isoforms. However, in vivo studies suggest that FGFR1c may be the most important for the activities of FGF21.
FGF21 mimetics are currently being studied in human trials to ascertain their utility in the treatment of obesity and type 2 diabetes. Studies with FGF21 mimetics in humans demonstrated that in obese individuals there was a significant improvement in overall serum lipid levels. FGF21 has also been shown to be involved in functions in the brain that result in reductions in the intoxicating effects of alcohol (ethanol) as well as reducing the desire to consume ethanol.
FGF21 Effects on Metabolism and Energy Homeostasis
FGF21 has been shown to be a critical integrator of various signals (hormonal, stress, etc.) that regulate metabolism. The effects of FGF21 include enhancing insulin sensitivity, increasing energy expenditure, decreasing hepatic triglyceride content, and regulation of overall macronutrient preference.
With respect to the effects of FGF21 on insulin sensitivity it has been shown to be one of the most potent acute sensitizers of insulin. For example, studies have demonstrated that a single injection of FGF21 into experimental animals with diet-induced obesity can result in up to a 50% reduction in plasma glucose levels. This effect of FGF21 is not observed in lean animals when FGF21 is injected alone, however, when co-injected with insulin the resultant plasma glucose disposal is greater than that seen with insulin alone.
FGF21 functions primarily by regulating glucose and lipid homeostasis during late fasting but also functions acutely during refeeding through its ability to enhance insulin-stimulated glucose uptake and suppression of lipolysis. The functions that have been shown to be exerted by FGF21 indicate that it serves as a metabolic switch to maintain nutrient homeostasis. In addition, FGF21 action sensitizes multiple endocrine signals during fasting, such as glucagon, and during refeeding, such as insulin. These effects allow for efficient metabolic transitions from the fasted to the refed state.
FGF21 Suppresses Ethanol Consumption
Induction of FGF21 expression in the liver occurs in response to numerous metabolic stresses including protein deficiency, starvation, and ethanol. Consumption and metabolism of ethanol results in the most potent increases in the production of FGF21 in the liver. The results of this increased FGF21 include central nervous system arousal and altered hepatic ethanol metabolism. The FGF21 produced by the liver enters the blood where it can then enter the brain and exert effects via binding to its receptor complex. The effects of FGF21 in the brain include a suppression of ethanol preference and an induction of water consumption. Polymorphisms in the FGF21 gene, as well as in the gene (KLB) encoding the FGF21 receptor binding partner, β-Klotho, have been found to be associated with a propensity for alcohol consumption.
Experiments in primates has shown that FGF21 exerts effects in the nucleus accumbens (NAc) by altering glutamate-mediated neural transmission. This, in turn, suppresses alcohol consumption through a projection-specific subpopulation of KLB-expressing neurons in the basolateral amygdala (BLA). Pharmacologic administration of an FGF21 analogue exerts the same effects as FGF21 released from the liver in response to normal physiological processes.
FGF21 Ameliorates Ethanol Intoxication
Within the brain, specifically within the locus coeruleus (LC), FGF21 directly activates the norepinephrine producing neurons. When norepinephrine is released from LC neurons it activates both α1– and β-adrenergic receptors in subcortical regions of the brain responsible for arousal. Experiments in mice have shown that the activation of the noradrenergic system of the LC, by FGF21, reduces the intoxication effects of ethanol. In this same animal model, knocking out the FGF21 gene leads to exacerbation of the intoxicating effects of ethanol. Conversely, the administration of FGF21 to mice following ethanol administration dramatically reduces the time required for recovery from the ethanol induced effects.
The ability of FGF21 to exert its anti-intoxicant effect can be blocked by interference with the adrenergic receptor system. Administration of either prazosin, a selective α1-adrenergic receptor blocker, and propranolol, a β-adrenergic receptor blocker, result in inhibition of the effects of FGF21 on the intoxicating effects of ethanol.
Human FGF Receptor-Related Disorders
Several other disorders of bone growth collectively identified as craniosynostosis syndromes have been shown to result from mutations in FGFR1, FGFR2 and FGFR3. Sometimes the same mutation can cause two or more different craniosynostosis syndromes. A cysteine to tyrosine substitution in FGFR2 can cause either Pfeiffer or Crouzon syndrome. This phenomenon indicates that additional factors are likely responsible for the different phenotypes.
Table of Human FGF Receptor-Related Disorders
| Affected Receptor | Syndrome | Phenotypes |
| FGFR1 | Pfeiffer | craniosynostosis (premature skull bone fusion); hypertelorism (wide-spaced eyes); micrognathia (small jaw); beaked nose; high forehead; brachydactyly (short fingers and toes); syndactyly (digit fusion); autosomal dominant inheritance; type 1 disease associated with either FGFR1 or FGFR2 mutations, type 2 and 3 disease caused by FGFR2 mutations only |
| FGFR2 | Apert | craniosynostosis (premature skull bone fusion); fusion of digits; mild to moderate intellectual impairment; autosomal dominant inheritance |
| FGFR2 | Beare-Stevenson | craniosynostosis (premature skull bone fusion); cutis gyrata (corrugated skin); acanthosis nigricans (thick, dark skin patches); fewer than 20 persons world-wide known to have disease; autosomal dominant inheritance |
| FGFR2 | Crouzon | craniosynostosis (premature skull bone fusion); ocular proptosis (eyeball protrusion); hearing loss; underdeveloped upper jaw; autosomal dominant inheritance |
| FGFR2 | Jackson-Weiss | craniosynostosis (premature skull bone fusion); syndactyly; wide and short big toes that point away from other toes; autosomal dominant inheritance |
| FGFR2 | Pfeiffer | same as for FGFR1 mutations |
| FGFR3 | Achondroplasia | short-limbed dwarfism; macrocephaly; lordosis (sway back); bowed legs; kyphosis (abnormal front-to-back spine curvature); autosomal dominant inheritance; 80% due to new mutations, highly correlated to increased age of father |
| FGFR3 | Crouzonodermoskeletal syndrome | craniosynostosis (premature skull bone fusion); acanthosis nigricans; beaked nose; underdeveloped upper jaw; ocular proptosis; strabismus (eyes that point in different directions); autosomal dominant inheritance |
Transforming Growth Factor-β (TGF-β)
A more detailed description of the TGF-β family of growth factors and associated signaling pathways can be found on the Signaling by Wnts and TGFs-β/BMP page.
TGF-β was originally characterized as a protein secreted from a tumor cell line that was capable of inducing a transformed phenotype in non-neoplastic cells in culture. This effect was reversible, as demonstrated by the reversion of the cells to a normal phenotype following removal of the TGF-β. Subsequently, many proteins homologous to TGF-β have been identified. The four closest relatives are TGF-β1 (the original TGF-β) through TGF-β5 (TGF-β1 is the same as TGF-β4). All four of these proteins share extensive regions of similarity in their amino acids. Many other proteins, possessing distinct biological functions, have stretches of amino-acid homology to the TGF-β family of proteins, particularly the C-terminal region of these proteins.
The TGF-β-related family of proteins includes the activin and inhibin proteins. There are activin A, B and AB proteins, as well as an inhibin A and inhibin B proteins. The Müllerian inhibiting substance (MIS) is also a TGF-β-related protein, as are members of the bone morphogenetic protein (BMP) family of bone growth-regulatory factors. Indeed, the TGF-β family may comprise as many as 100 distinct proteins, all with at least one region of amino-acid sequence homology.
There are several classes of cell-surface receptors that bind different TGFs-β with differing affinities. There also are cell-type specific differences in receptor sub-types. Unlike the EGF, PDGF and FGF receptors, the TGF-β family of receptors all have intrinsic serine/threonine kinase activity and, therefore, induce distinct cascades of signal transduction.
TGFs-β have proliferative effects on many mesenchymal and epithelial cell types. Under certain conditions TGFs-β will demonstrate anti-proliferative effects on endothelial cells, macrophages, and T- and B-lymphocytes. Such effects include decreasing the secretion of immunoglobulin and suppressing hematopoiesis, myogenesis, adipogenesis and adrenal steroidogenesis. Several members of the TGF-β family are potent inducers of mesodermal differentiation in early embryos, in particular TGF-β and activin A.
Transforming Growth Factor-α (TGF-α)
TGF-α, like the original founding member of the TGF-β family, was first identified as a substance secreted from certain tumor cells that, in conjunction with TGF-β1, could reversibly transform certain types of normal cells in culture. The TGF-α precursor protein is derived from the TGFA gene which is located on chromosome 2p13 and is composed of 7 exons that generate four alternatively spliced mRNAs. The predominant TGF-α preproprotein contains 160 amino acids.
Following processing of the preproprotein, TGF-α can be function as a transmembrane-bound ligand or it can be fully processed to the secreted extracellular growth factor form. TGF-α binds to the EGF receptor and it is this interaction that is responsible for the growth factor’s effect.
The predominant sources of TGF-α are carcinomas, but activated macrophages, keratinocytes (and possibly other epithelial cells), and hypothalamic astrocytes also produce and secrete TGF-α. In normal cell populations, TGF-α is a potent keratinocyte growth factor; forming an autocrine growth loop by virtue of the protein activating the very cells that produce it. Within the brain, TGF-α regulates the synthesis and release of the anterior pituitary hormone, luteinizing hormone-releasing hormone, LHRH. This latter effect of TGF-α is important in the maturation of the secondary female sex characteristics.
Erythropoietin (EPO)
Erythropoietin (EPO) is a glycoprotein hormone that is synthesized by interstitial fibroblasts near the proximal convoluted tubules of the kidney. Erythropoietin is the primary regulator of erythropoiesis. Although EPO is synthesized by the fetal liver, this location is of limited significance to EPO synthesis in the adult. Synthesis and release of EPO occurs in response to hypoxic conditions. EPO stimulates the proliferation and differentiation of immature erythrocytes; it also stimulates the growth of erythroid progenitor cells (e.g. erythrocyte burst-forming and colony-forming units: CFU-E) and induces the differentiation of erythrocyte CFU-E into proerythroblasts.
The EPO protein is derived from the EPO gene which is located on chromosome 7q22.1 and is composed of 5 exons that encode a 193 amino acid precursor protein.
The effects of EPO are exerted in response to the hormone binding to a specific EPO receptor. Activation of the EPO receptor results in signal transduction events involving the JAK/STAT pathway. The EPO receptor is derived from the EPOR gene which is located on chromosome 19p13.2 and is composed of 8 exons that encode a 508 amino acid precursor protein.
When patients suffering from anemia, due to kidney failure or as a result of cancer therapy, are administered human recombinant EPO (hrEPO: Procrit and Epogen), the result is a rapid and significant increase in red blood cell count.
Insulin-Like Growth Factor-1 (IGF-1)
IGF-1 (originally called somatomedin C) is a growth factor structurally related to insulin. IGF-1 is the primary protein involved in responses of cells to growth hormone (GH): that is, IGF-1 is produced in response to GH and then induces subsequent cellular activities. It is the activity of IGF-1, in response to GH, that gave rise to the term somatomedin. Subsequent studies demonstrated that IGF-1 has autocrine and paracrine activities in addition to the initially observed endocrine activities on bone. IGF-1 belongs to the insulin-like growth factor system that includes IGF-1, IGF-2 (described in the next section), IGF binding proteins, and the receptors that bind the growth factors.
The IGF-1 precursor is derived from the IGF1 gene which is located on chromosome 12q23.2 and is composed of 7 exons that generate multiple mRNAs via alternative splicing and alternative polyadenylation site utilization. In addition, IGF1 gene expression is controlled by multiple transcriptional initiation sites. Two classes of IGF-1 mRNA result from this complex control such that class 1 mRNAs initiate from promoter elements in exon 1, whereas class 2 mRNAs initiate from promoters in exon 2. IGF-1 mRNAs initiating from promoter 1 in exon 1 are found in multiple tissues, whereas, transcriptional initiation from promoter 2 in exon 2 is restricted to the liver and the kidney. Expression of the IGF-1 gene in the liver is the major source of secreted IGF-1 hormone accounting for 75% of total serum IGF-1. The longest IGF-1 preproprotein contains 158 amino acids. Despite the complex transcriptional regulation and the generation of multiple prepro-IGF-1 proteins, all of the resultant mature hormones are 70 amino acids in length.
IGF-1 exerts its biological effects primarily as a result of binding to, and activating, the IGF-1 receptor. The IGF-1 receptor (IGF1R), like the insulin receptor, is composed of disulfide bonded α- and β-peptides that are derived by proteolytic processing of the primary translation product. In addition, like the insulin receptor, the IGF1R has intrinsic tyrosine kinase activity. Owing to their structural similarities IGF-1 can bind to the insulin receptor but does so at a much lower affinity than does insulin itself. The IGF1R gene is located on chromosome 15q26.3 and is composed of 24 exons that generate two alternatively spliced mRNAs encoding IGF1R isoform 1 precursor (1367 amino acids) and isoform 2 precursor (1366 amino acids).
In addition to binding to the IGF-1 receptor, IGF-1 activity (as well as IGF-2 activity: next section) is controlled by binding to one of several IGF binding proteins (IGFBP). Humans express six IGFBPs (IGFBP1–IGFBP6) that sequester IGFs in serum resulting in control of their interaction with IGF receptors. About 75% of circulating IGFs are bound in ternary complexes that are composed of IGF-1 or IGF-2, IGFBP-3 and IGFBP acid-labile subunit (IGFALS). IGFALS is synthesized by, and secreted from, the liver. The IGFBP1 gene is located on chromosome 7p12.3 and is composed of 4 exons that encode a 259 amino acid precursor protein. The IGFBP2 gene is located on chromosome 2q35 and is composed of 4 exons that generate four alternatively spliced mRNAs encoding three distinct precursor proteins, only one of which is secreted. The IGFBP3 gene is located on chromosome 7p12.3 and is composed of 5 exons that generate two alternatively spliced mRNAs encoding isoform a precursor (297 amino acids) and isoform b precursor (291 amino acids). The IGFBP4 gene is located on chromosome 17q21.2 and is composed of 4 exons that encode a 258 amino acid precursor protein. The IGFBP5 gene is located on chromosome 2q35 and is composed of 4 exons that encode a 272 amino acid precursor protein. The IGFBP6 gene is located on chromosome 12q13 and is composed of 4 exons that encode a 240 amino acid precursor protein. The IGFALS gene is located on chromosome 16p13.3 and is composed of 4 exons that generate two alternatively spliced mRNAs encoding two distinct precursor proteins.
Insulin-Like Growth Factor-2 (IGF-2)
Like IGF-1, IGF-2 is a growth factor that is structurally related to insulin and shares 67% amino acid identity with IGF-1. IGF-2 is almost exclusively expressed in embryonic (fetal and placental) and neonatal tissues. Following birth, in humans, the level of IGF-2 rises in early childhood and remains relatively steady throughout adulthood until old age when it declines. During adult life the level of serum IGF-2 is approximately 3 times that of IGF-1. Due to the fetal and placental expression of IGF-2, it was originally thought to be primarily a fetal growth factor. However, evidence clearly indicates that IGF-2 does indeed exert important metabolic effects in the adult. The primary tissue responsible for adult IGF-2 expression is the liver. The IGF2 gene is located on chromosome 11p15.5 and is composed of 9 exons that generate five alternatively spliced mRNAs that collectively encode two distinct preproproteins of 236 and 180 amino acids. The 236 amino acid preproprotein originates from translational initiation at an upstream AUG codon not found in two of the other four mRNA splice variants. Prepro-IGF-2 contains a 24-amino acid signal peptide. Within the Golgi apparatus, pro-IGF-2 is O-glycosylated and then further proteolyzed to the mature IGF-2 form via the action of proprotein convertase subtilisin/kexin type 4 (PCSK4). PCSK4 is historically referred to as prohormone convertase 4 (PC4). Post-translational processing of IGF-2 is an incomplete process such that several pro-IGF-2 peptides (collectively referred to as “big” IGF-2) are secreted into the blood, accounting for 10%–20% of total serum IGF-2.
Expression of the IGF2 gene is controlled by the epigenetic phenomenon of genomic imprinting. Expression of the IGF2 gene is restricted to the paternal allele, in most tissues, via the imprinting phenomenon. The IGF-2 gene also contains four promoters (identified as P1–P4) from which IGF-2 is transcribed. The P2–P4 promoters control IGF-2 transcription in the embryo, whereas transcription occurs from all four promoters in the liver of adult humans. Expression of the IGF-2 gene in the liver of adults occurs from both the paternal and the maternal alleles which may explain why circulating IGF-2 concentrations remain elevated throughout adult life.
IGF-2 exerts its biological effects by interacting with the IGF1R as well as the A form of the insulin receptor (IR-A). IGF-2 also binds to another receptor, that is specific for this particular IGF family member, identified as the IGF2R. The IGF2R is also a mannose-6-phosphate (M6P) receptor, similar to the M6P receptor that is responsible for the integration of lysosomal enzymes (which contain mannose-6-phosphate residues) into the lysosomes. Binding of IGF-2 to the IGF2R is responsible for clearance of IGF-2 from the circulation and does not contribute to IGF-2-mediated signal transduction. The IGF2R gene is located on chromosome 6q26 and is composed of 48 exons that encode a 2491 amino acid precursor protein. The IGF2R protein is a cation-independent mannose-6-phosphate receptor and therefore, is also referred to as the M6P/IGF2 receptor. In addition to IGF-2, the IGF2R has been shown to bind a diverse array of mannose-6-phosphate-containing proteins as well as several non-glycosylated proteins.
The initial observations that suggested the role of IGF-2 was most significant for fetal development only were obtained in knockout mouse studies. In these mice, fetal development was severely retarded yet following birth the mice grew normally and were fertile. Whereas in mice the level of IGF-2 falls following birth, in humans it does not. Studies in humans have shown that IGF-2 has a role in fetal growth and development by promoted formation of mesodermal germ layer. The level of IGF-2, in utero, is ten times higher than that of IGF-I. The growth effects of IGF-2 during fetal development are primarily exerted by its binding to, and activating, the IR-A form of the insulin receptor. Fetal actions of IGF-2 also involve activation of the IGF1R. In addition to its role in fetal development, IGF-2 is also an important regulator of placental growth where it promotes nutrient transport, trophoblast invasion and proliferation and survival of cytotrophoblasts.
Postnatally IGF-2 exerts a potent angiogenic effect, central to its role in organ development and maintenance. The angiogenic effect of IGF-2 is the result of the growth factor inducing an up regulation of the expression of vascular endothelial growth factor, VEGF. IGF-2 has also been shown to exert growth promoting effects within the immune system. IGF-2 promotes granulocyte macrophage colony formation, stimulates the growth of B cells, and stimulates the growth of erythroid and myeloid precursor cells. Within specific organ systems IGF-2 exerts import growth and proliferative effects such as pancreatic β-cell proliferation and survival, development and maintenance of the musculoskeletal system, and development of bone. IGF-2, like insulin, exerts both growth factor functions and metabolic hormone regulating effects. These hormonal effects are most significant within adipose tissue, skeletal muscle and liver. Within the liver, IGF-2 actions result in the suppression of hepatic glucose output and increased glycogen synthesis. In adipose tissue and skeletal muscle, as well as several other peripheral tissues, IGF-2 induces glucose uptake and oxidation and increases synthesis of lipids and proteins.
In addition to its normal growth and metabolic functions, dysregulation of IGF-2 function has been associated with numerous pathologies, in particular in obesity and type 2 diabetes. Serum IGF-2 concentrations have been shown to increase in obesity and these levels correlate positively with BMI. The increased serum concentration of IGF-2 in obesity most likely represents an appropriate physiological response designed to promote energy storage in response to increased dietary supply. When overweight and obese individuals lose weight there is an associated decrease in total serum IGF-2 levels. In humans with normally low levels of serum IGF-2 there is an increased risk for weight gain and obesity. Although the mechanism by which the low levels of IGF-2 contribute to future weight gain is not clearly understood, it is an important prognostic indicator.
Numerous studies have shown that IGF-2 is also dysregulated in diabetes. In type 2 diabetics, who are also obese, the levels of IGF-2 are even higher than in obese individuals that do not also exhibit insulin resistance typical of type 2 diabetes. Although the cause of the diabetes-related increase in IGF-2 is unknown, it believed to be primarily the result of increased adipose tissue secretion in response to hyperglycemia. The increases in serum IGF-2 seen in obese individuals has been shown to predispose these individuals to future development of insulin resistance and type 2 diabetes.
Several studies have shown a strong correlation between obesity and cancer. In this context, the IGF systems are known to be causally linked to this phenomenon. The contribution to cancer development in obese patients, where IGF-2 levels are elevated, is thought to be exerted via IGF-2 binding to the IR-A form of the insulin receptor. When IGF-2 binds and activates this receptor, a set of signaling proteins, distinct from those activated by insulin binding, is activated that favor a mitogenic program increasing the likelihood for cancer.
Obesity in pregnant females has been shown to correlate with epigenetic changes in the IGF2 gene. These changes are reflected in a reduced level of methylation of the control region of the maternal IGF2 gene leading to increased expression of IGF-2 and increased IGF-2 concentrations in umbilical cord blood. The consequences of the altered maternal epigenome are evidenced by an adverse metabolic health of the fetus. Paternal obesity has also been associated with changes in the epigenome of the IGF2 gene. However, in the case of paternal obesity the reduced methylation is observed in the fetal IGF2 gene. Defects in imprinting at the IGF2 locus are seen in Beckwith-Wiedemann syndrome, BWS. As a result of the chromosomal alterations in BWS patients there is fetal overgrowth, organomegaly and an increased risk of developing tumors. Dysregulated, over-expression of IGF-2 in the BWS fetus is believed to account for the majority of the clinical features observed in this disease.
Tumor Necrosis Factor (TNF) Superfamily
The tumor necrosis factor superfamily (TNFSF) of signaling proteins, and the receptors to which these proteins bind (identified as the TNF receptor superfamily, TNFRSF), represents an important class of proteins involved in the regulation of a variety of stimulus-responsive processes. Humans express 18 genes that encode proteins of the TNF superfamily. Although the TNFSF proteins are known to be required for numerous processes in cells, many of the proteins are critical to the regulation of immune functions. The original member of the TNFSF is tumor necrosis factor which is encoded by the TNF gene. At one point this protein was called TNF alpha (TNFα; as well as cachectin). Proteins of the TNFSF are major immune response-modifying cytokines produced primarily by activated macrophages. TNF induces the expression of other autocrine growth factors, increases cellular responsiveness to growth factors and induces signaling pathways that lead to proliferation. Like other growth factors, TNF induces expression of a number of nuclear proto-oncogenes as well as several interleukins and pro-inflammatory cytokines.
Table of Human TNFSF Proteins
| TNFSF Gene Symbol | Protein Name(s) | Functions / Comments |
| TNF | tumor necrosis factor-alpha (TNFα); cachectin | component of acute phase reaction of inflammatory responses; induces fever, activates apoptosis, inhibits tumorigenesis, inhibits viral replication; synthesized primarily by macrophages but also by other leukocytes such as neutrophils, mast cells, eosinophils, and natural killer (NK) cells; exerts its effects by binding to two TNF receptor superfamily (TNFRSF) members: TNFR1 (encoded by the TNFRSF1A gene) and TNFR2 (encoded by the TNFRSF1B gene) |
| LTA | lymphotoxin-α (LT-α); tumor necrosis factor-beta; TNFβ | secreted and forms a homotrimeric complex; kills a number of different cell types; induces terminal differentiation in many cell types; inhibition of lipoprotein lipase (LPL) present on the surface of vascular endothelial cells; can form heterodimers with LT-β; predominant site of synthesis is cytotoxic T-lymphocytes (CTL); binds to TNFR1, TNFR2, and HVEM (Herpes Virus Entry Mediator; encoded by the TNFRSF14 gene) |
| LTB | lymphotoxin-β (LT-β); tumor necrosis factor C (TNF-C) | membrane-associated (type II transmembrane) protein; LT-α binds to cell-surface LT-β in a LT-α1/LT-β2 heterotrimeric complex; the LT-α1/LT-β2 complex induces inflammatory responses through its binding to the LT-β receptor (encoded by the TLTBR gene) on adjacent cells; the LTBR encoded protein is a member of the TNF receptor superfamily identified as TNFRSF3 |
| TNFSF4 | TNF superfamily member 4; OX40 ligand | TNFSF4 binds to the cell surface CD134 protein; enhances Th2 type cell differentiation; binds to the cell surface OX40 protein which is a member of the TNF receptor superfamily (encoded by the TNFRSF4 gene) |
| CD40LG | CD40 ligand; TNF superfamily member 5 | expressed predominantly on activated T cells; binds to the cell surface CD40 protein (also identified as TNFRSF5) on antigen presenting cells |
| FASLG | FAS ligand; FASL; TNF superfamily member 6 | membrane-associated (type II transmembrane) protein which contains death domain (DD) in intracellular portion; functions as a homotrimeric complex by interacting with the cell surface receptor identified as FAS; interaction of FASL on one cell with FAS on an adjacent cell triggers apoptosis via the formation of what is referred to as a death-inducing signaling complex (DISC); upon FASL:FAS interaction the intracellular adapter protein FAS-associated death domain (FADD) binds to the DD of FASL triggering the signaling resulting in activation of apoptosis; in the TNF receptor superfamily nomenclature FAS is TNFRSF6 |
| CD70 | TNF superfamily member 7; CD27 ligand | expressed on activated T and B cells, particularly T and B lymphoma cells; binds to cell surface CD27 (also identified as TNFRSF7) |
| TNFSF8 | TNF superfamily member 8; CD30 ligand | binds to cell surface CD30 (encoded by the TNFRSF8 gene) which which is a member of the TNF receptor superfamily |
| TNFSF9 | TNF superfamily member 9; 4-1BB ligand | membrane-associated (type II transmembrane) protein; expressed on activated T cells; binds to the TNF receptor superfamily protein encode by the TNFRSF9 gene also identified as 4-1BB |
| TNFSF10 | TNF superfamily member 10; TNF-Related Apoptosis-Inducing Ligand (TRAIL) | expressed and secreted by numerous cell types; induces apoptosis in a variety of tumor cells; binds to several TNF receptor superfamily proteins that are members of the death receptor (DR) subfamily; binds to several transmembrane and soluble proteins including TRAIL receptor 1 (TRAILR1, TNFRSF10A, DR4), TRAILR2 (TNFRSF10B, DR5), TRAILR3 (this is a soluble decoy receptor also known as DcR1; encoded by the TNFRSF10C gene), TRAILR4 (TNFRSF10D, DcR2), and TNFRSF11B (also known as osteoprotegrin: OPG) |
| TNFSF11 | TNF superfamily member 11; Receptor Activator of NF-κB Ligand (RANKL) | membrane-associated (type II transmembrane) protein; regulates apoptosis and also controls bone regeneration and remodeling; binds to RANK (encoded by the TNFRSF11A gene) |
| TNFSF12 | TNF superfamily member 12; TNF-related WEAK inducer of apoptosis (TWEAK) | activity similar to that of TNF; binds to the TWEAK receptor encoded by the TNFRSF12A gene |
| TNFSF13 | TNF superfamily member 13; A PRoliferation-Inducing Ligand (APRIL) | involved in B cell development; functions by binding to the TNF receptor superfamily 17 protein encoded by the TNFRSF17 gene (also identified as B cell maturation antigen, BCMA) |
| TNFSF13B | TNF superfamily member 13B; B-cell Activating Factor (BAFF) | membrane-associated (type II transmembrane) protein; expressed on numerous cell types; also known as B lymphocyte stimulator (BLyS); activates NF-kB signaling; binds three TNF receptor superfamily members encoded by the TNFRSF13B (also known as transmembrane activator and calcium modulator and cyclophilin ligand interactor, TACI), TNFRSF13C (also known as BAFF receptor, BAFFR), and TNFRSF17 |
| TNFSF14 | TNF superfamily member 14; LIGHT (homologous to Lymphotoxin, exhibits Inducible expression and competes with HSV Glycoprotein D for binding to Herpesvirus entry mediator, a receptor expressed on T lymphocytes) | secreted protein; functions as a costimulatory molecule and an inhibitor to herpesvirus infection; binds to the TNF receptor superfamily proteins encoded by the TNFRSF14 (also known as HVEM) and TNFRSF6B (also identified as decoy receptor 3, DcR3) genes |
| TNFSF15 | TNF superfamily member 15; Vascular Endothelial cell Growth Inhibitor (VEGI) | abundantly expressed by endothelial cells; expression induced TNF and IL-1α; binds to the TNF receptor superfamily 25 protein (encoded by the TNFRSF25 gene) which is also identified as death receptor 3 (DR3); functions as an autocrine inducer of apoptosis in endothelial cells |
| TNFSF18 | TNF superfamily member 18; Glucocorticoid-Induced TNF Receptor family-related gene Ligand (GITRL) | expressed in endothelial cells; binds to the TNF receptor superfamily 18 protein encoded by the TNFRSF18 gene |
| EDA | ectodysplasin A | membrane-associated (type II transmembrane) protein; functional as a homotrimeric complex; involved in the development of ectodermal organs; binds to the ectodysplasin A receptor (encoded by the EDAR gene) |
TNF Receptor Super Family: TNFRSF
The TNF receptor superfamily (TNFRSF) members are transmembrane proteins having cysteine-rich motifs in their extracellular domains. Humans express 29 genes that encode receptor proteins of the TNF receptor superfamily. Many of the genes that express receptors for TNF superfamily member proteins are identified by the designation TNFRSF followed by a number. For example the TNF receptors identified as TNFR1 and TNFR2 are encoded by the TNFSFR1A and TNFRSF1B genes, respectively.
The 29 TNFRSF proteins can be divided into three different subfamilies that are dependent upon the specific intracellular signals that are induced by the receptors upon ligand binding. These three receptor groups are the death domain (DD)-containing receptors, the decoy receptors (DcR), and the TNF receptor-associated factor (TRAF)-binding receptors.
The DD-containing members of the TNFRSF include TNFR1 (TNFRSF1A), FAS (TNFRSF6), death receptor 3 (DR3; encoded by the TNFRSF25 gene), DR4 (TNFRSF10A), DR5 (TNFRSF10B), and DR6 (TNFRSF21).
The TNF receptor superfamily proteins that contain a TRAF-interacting motif (TIM) include TNFR2 (TNFRSF1B), lymphotoxin-β receptor (LTBR gene; also identified as TNFRSF4), OX40 (TNFRSF4), CD40 (TNFRSF5), CD27 (TNFRSF7), CD30 (TNFRSF8), receptor activator of NF-κB (RANK; encoded by the TNFRSF11A gene), and glucocorticoid-induced TNF receptor family-related gene (GITR; encoded by the TNFRSF18 gene).
As the name implies, the TRAF-binding subfamily of TNFRSF proteins bind TRAF proteins which are adaptor molecules that activate multiple downstream signaling pathways such as NF-κB, Janus kinase (JNK), ERK, p38MAPK, and PI3K.
Although some TNFRSF proteins contain their own intracellular death domains, other members interact with intracellular proteins that contain death domains. These intracellular DD proteins serve as scaffolds for binding pro-caspases which then undergo auto-cleavage. The cleavage of a pro-caspase to an active caspase results in the formation of the death-inducing signaling complex (DISC) and induction of apoptosis.
Clinical Utility of TNFSF Inhibition
Inhibition of the activity of members of the TNF superfamily has proven to be effective in the treatment of certain autoimmune disorders such as rheumatoid arthritis, RA. Currently there are seven anti-TNF biologicals (Adalimumab, Adalimumab-atto, Etanercept, Etanecept-szzs, Infliximab, Golimumab, and Certolizumab) that have been approved for use by the US FDA. These therapies are used in the treatment of various forms of autoimmune disease such as RA, psoriatic arthritis, ulcerative colitis, Crohn disease, ankylosing spondylitis, and juvenile arthritis. Several of these biologicals bind to soluble TNF molecules and prevent their binding to their cognate receptors resulting in a loss of the normal activation of synthesis and secretion of pro-inflammatory cytokines such as IL-1 and IFN-γ.
Hedgehog Family Factors
The original hedgehog protein was identified in the fruit fly, Drosophila melanogaster, because mutant fruit fly larva demonstrated an abnormal pattern of spiky projections, called denticles, on their exoskeletons that resembled the spines of a hedgehog.
Humans express three members of the hedgehog family of signaling molecules identified as Indian hedgehog (Ihh), desert hedgehog (Dhh), and sonic hedgehog (Shh). These proteins are important for the organogenesis of almost all organs as well as being involved in organ regeneration, and organ homeostasis.
Mutations, deficiencies, and abnormal levels of expression of hedgehog proteins, as well as other genes in the hedgehog signaling pathways lead to an array of disorders. These disorders include malformations, hyperplasia, and growth retardation within the central nervous system (CNS) as well as in the processes of craniofacial, limb, and skeletal developmental. On the other side of the spectrum, the persistent or aberrantly high expression and function of hedgehog proteins is a significant contributor to tumorigenesis in organs at postnatal stages.
Indian hedgehog is encoded by the IHH gene. The IHH gene is located on chromosome 2q35 and is composed of 3 exons that encode a preproprotein of 411 amino acids.
Desert hedgehog is encoded by the DHH gene. The DHH gene is located on chromosome 12q13.12 and is composed of 5 exons that encode a preproprotein of 396 amino acids. The highest levels of DHH expression are found in the testes.
Sonic hedgehog is encoded by the SHH gene. The SHH gene is located on chromosome 7q36.3 and is composed of 7 exons that generate two alternatively spliced mRNAs. These mRNAs encode preproproteins of 462 amino acids (isoform 1) and 179 amino acids (isoform 2).
Sonic Hedgehog, Shh
All hedgehog proteins undergo the same processing steps in the endoplasmic reticulum (ER) and Golgi apparatus, but only the details of Shh will be addressed in this section.
Following the synthesis of the Shh preproprotein in the endoplasmic reticulum (ER) it undergoes auto proteolytically cleavage into two secreted peptides. These peptides are 19 kDa (amino terminus) and 26 kDa (carboxy terminus). The 19 kDa peptide is identified as Shh-N while the 26 kDa peptide is identified as Shh-C.
The Shh-N peptide mediates Shh signaling while the the Shh-C peptide is responsible for regulation of the auto proteolysis reaction. During the proteolysis a cholesterol moiety is added at the C-terminus of Shh-N via the cholesterol transferase activity associated with the C-terminal ends of the Shh peptides. Following the addition of the cholesterol the Shh-C peptide diffuses away and a fatty acid (primarily palmitic acid) is added to the N-terminus of the Shh-N peptide. The fatty acylation of Shh-N (primarily palmitoylation) is catalyzed by an acyltransferase that was initially referred to as Skinny hedgehog (Ski) acyltransferase but is now known as hedgehog acyltransferase (encoded by the HHAT gene). The addition of cholesterol is critical for the secretory regulation and long-range activity of Shh while the palmitoylation increases the inductive potency of Shh.
The HHAT encoded acyltransferase belongs to a family of enzymes identified as the membrane bound O-acyltransferase (MBOAT) family. The MBOAT family of transferases includes several enzymes involved in the synthesis of triglycerides and in the synthesis of phospholipids.
The hedgehog acyltransferase is localized to the membranes of the endoplasmic reticulum. The mobilization of cholesterol from the endosomes into the ER is the function of the Niemann-Pick proteins NPC1 and NPC2. The ER membrane-associated protein called dispatched (Disp) and the secreted protein Scube2 (signal peptide, CUB domain and EGF like domain containing 2) bind to the cholesterol moiety of the modified Shh protein which promotes release from the cell membrane. Humans express three genes encoding dispatched proteins, DISP1, DISP2, and DISP3. These genes are officially referred to as dispatched RND transporter family genes where RND refers to resistance-nodulation-division transporters identified in Gram-negative bacteria. Humans express three genes that encode Scube proteins, SCUBE1, SCUBE2, and SCUBE3.
Shh Signaling
Following the release of Shh from its cell of synthesis it will bind to target cells via the interaction a specific plasma membrane receptor. The primary receptor for Shh is identified as Patched (Ptch). Patched is a protein that spans the plasma membrane 12 times and is localized to ciliary membranes in the absence of Shh. Following the binding of Shh the Shh-Ptch complex moves to the plasma membrane. The Shh-Ptch complex is then endocytosed, a process that requires the membrane-remodeling GTPase called dynamin and the HECT-domain ubiquitin E3 ligases known as Smurf1/2. The incorporation of the Shh-Ptch complexes to the endosome results in their degradation.
The removal of Ptch from the ciliary membrane allows for the GPCR identified as smoothened (Smo) to translocated from the plasma membrane to the ciliary membrane. While Smo is mobilized to the ciliary membrane it is phosphorylated on Ser residues in the intracellular C-terminal domain by two kinases, casein kinase 1α (CK1α) and GPCR kinase 2 (GRK2). Phosphorylated Smo then interacts with β-arrestin 2. The interaction of Smo with β-arrestin 2 regulates the activity and the stability of the Smo protein.
Smo activates the translocation of the transcription factors Gli2 and Gli3 (often designated Gli2/3) to the nucleus. During the mobilization of Gli2/3 to the nucleus they are phosphorylated, acetylated, and SUMOylated. The kinases that phosphorylate Gli2/3 are PKA and glycogen synthase kinase 3 (GSK3).
Gli family transcription factors are Krüppel-like transcription factors that contain a zinc-finger DNA-binding domain. The Gli proteins are dual activity transcription factors where the N-terminus contains a transcriptional repressor domain and the C-terminus contains an activator domain. In the absence of Shh signaling Gli2/3 are proteolyzed and the transcription repressor N-terminal domain is translocated into the nucleus and represses the transcription of target genes. In the presence of Shh signaling the Gli2/3 proteins are not cleaved and are the substrates for phosphorylation, acetylation, and SUMOylation.
Within the nucleus, Gli2/3 binds to the target sequence, GACCACCCA/TGGGTGGTC. Shh target genes include those encoding Ptch and Gli1 as well as those encoding Nkx6.1, Olig2, Nkx2.2, and FoxA2.